Ecology conference in Iceland - prized students
Last Friday the Annual ecology conference in Iceland took place, in Keldnaholt Reykjavík. Four of our students presented their studies.
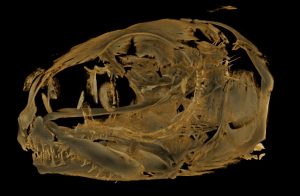
Ruhila Goswami gave her poster Diversity in craniofacial elements by populations in salmonids of Iceland. She won best student poster.
Andreas talked about Ants in Icelandic Geothermal Areas. He is wrapping up his M.Sc. thesis these weeks and will graduate in October 2025.
Kenedy Williams reported on her analyses of Feeding Element Specialization in Polymorphic Arctic Charr. She won an explorer grand from the Icelandic Biological society some weeks ago, for project on where the charr go when rivers warm.
Guðbjörg presented her new thesis chapter, on Divergence of functional phenotypes in feeding elements of Arctic charr (Salvelinus alpinus) along the benthic – pelagic axis. She won the best student talk prize.
Sex is not binary
Wrote couple of pieces for the icelandic web of science, a Q and A web about scientific topics, about sex related traits and their diversity in nature.
They are in Icelandic, but the central idea is that sex is quite variable in nature. Even in humans a range of traits are found in sex related structures, behaviors and perceptions.
All of it is quite natural. See for instance. Arnar Pálsson. „Hvað hefur áhrif á kyneinkenni og kynhneigð?“ Vísindavefurinn, 5. febrúar 2025, sótt 1. apríl 2025, https://visindavefur.is/svar.php?id=87491.
Silja Bára for rector
Wrote a letter of support for the candidacy of Silja Bara Omarsdottir for rector of the University of Iceland.
On icelandic news website Visir.is Mannlegi rektorinn Silja Bára - Vísir
Antteam visiting high schools
These weeks the Antteam at the University of Iceland is visiting high schools in Reykjavík and neighboring towns.
We have already been to FB, FG, MH, Kvennaskólinn and Verslunarskólinn, with FÁ scheduled this Wednesday.
Our graduate student Andreas Guðmundsson and Marco Mancini, along with Rafn Sigurðsson, doing an MSc on invasive mosses, are the main team members.
See more on our ants in iceland web page, maurar.hi.is.
Submission: Whole genome sequencing reveals how plasticity and genetic differentiation underlie sympatric morphs of Arctic charr
Whole genome sequencing reveals how plasticity and genetic differentiation underlie sympatric morphs of Arctic charr
Khrystyna Kurta , Mariano Olivera Fedi , and many others. Submitted to Nature Ecology and Evolution, and then Nature Communications.
Salmonids have a remarkable ability to form sympatric morphs after postglacial colonization of freshwater lakes. These morphs often exhibit differences in morphology, feeding, and spawning behaviour. Here we explore the genetics of morph differentiation by establishing a high-quality, annotated reference genome for the Arctic charr and use this as a resource for population genomic analysis of morphs from two Norwegian and two Icelandic lakes. The four lakes represent the spectrum of genetic differentiation between morphs from one lake with no genetic differentiation between morphs, implying phenotypic plasticity only, to two lakes with locus-specific genetic differentiation, implying incomplete reproductive isolation, and one lake with strong genome-wide divergence consistent with complete reproductive isolation. As many as 12 putative inversions ranging from 0.45 to 3.25 Mbp in size segregated among the four morphs present in one lake, Thingvallavatn, and these contributed significantly to the genetic differentiation among morphs. None of the putative inversions was found in any of the other lakes, but there were cases of partial haplotype sharing in similar morph contrasts in other lakes. The results are consistent with a highly polygenic basis of morph differentiation with limited genetic parallelism between lakes. The results support a model where morph differentiation is usually first established due to phenotypic plasticity that results in niche expansion and separation, followed by gradual development of reproductive isolation and locus-specific differentiation and eventually complete reproductive isolation and genome-wide divergence. A major explanation for salmonids’ ability to diversify into multiple sympatric morphs is likely the genome complexity caused by their ancient whole genome duplication that enhances evolvability.
Preprint: Gene flow and habitat heterogeneity shape coexistence dynamics of Arctic charr morphs in connected lakes
Han Xiao, Arnar Palsson, Zophonías Oddur Jónsson, et al. Gene flow and habitat heterogeneity shape coexistence dynamics of Arctic charr morphs in connected lakes. Authorea. December 27, 2024. DOI: 10.22541/au.173531304.47377188/v1
Submitted to Molecular Ecology.
Abstract
Preprint: Variation of tooth traits in ecologically specialized and sympatric morphs
Variation of tooth traits in ecologically specialized and sympatric morphs
Guðbjörg Ósk Jónsdóttir*, Finnur Ingimarsson, Sigurður Sveinn Snorrason, Sarah Elizabeth Steele, Arnar Pálsson. Submitted to Evolutionary Biology, link to bioRxiv.
GOJ: https://orcid.org/0009-0008-0502-5553
FI: https://orcid.org/0000-0002-0815-7622
SES: https://orcid.org/0000-0001-8404-5537
AP: https://orcid.org/0000-0002-6525-8112
Abstract
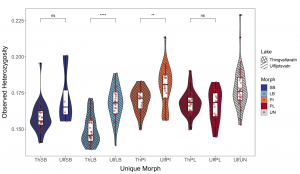
Differences in dentition between species relate to feeding specialisations, as examples of tetrapod dentition variation show clearly. The association of tooth traits and specialisations in non-mammalian vertebrates is less studied. We examined variation in dental traits in four sympatric morphs of Arctic charr (Salvelinus alpinus) which differ in feeding specialisations, head and jaw bone morphology. We studied tooth numbers in six bones (dentary, maxilla, premaxilla, palatine, vomer and glossohyal) and tooth angles in one bone (maxilla). We found fluctuating asymmetry in tooth numbers and angles and that the allometry of tooth numbers varied by bone but not morphs. The tooth numbers differed by morphs in four bones (dentary, palatine, vomer and glossohyal). In general, the morphs defined as pelagic had more teeth, and this relates partially to changes in bone shape. There was a difference in maxilla tooth angle, with benthic morphs having teeth which were angled more inwards. Dentary and maxilla tooth number correlated moderately with bone shape, maxilla tooth angle and premaxilla tooth number did not. While it is currently unknown what tooth characteristics are ancestral vs derived in these populations, the marked differences in specific bones presents an opportunity to explore rapid adaptive evolution in dentition.
Uppeldissvæði laxfiska í Þingvallavatni og tengdum ám
Now published in the Icelandic Naturalist. Areas used by Salmonid juveniles in Lake Þingvallavatn and connected rivers. Uppeldissvæði laxfiska í Þingvallavatni og tengdum ám (in icelandic, w. English abstract). Guðbjörg Ósk Jónsdóttir, Benóný Jónsson, Magnús Jóhannsson og Arnar Pálsson. (Náttúrufræðingurinn).
Nursery grounds of juvenile fish provide them shelter and food making them essential for the life of an individual, the population and species as a whole. In Lake Þingvallavatn, Iceland, two species of salmonids inhabit the lake, Brown trout (Salmo trutta) and Arctic charr (Salvelinus alpinus). Extensive biological studies have been conducted on adult fish of both species in the lake and its catchment area, but much less is known about the biology of age-0 fish and the ecology of their first 1−2 years in the life. Given the critical importance of nursery areas for the existence of fish populations and their contribution to biological diversity, research was undertaken to examine the distribution of charr and trout juveniles in the surf zone of Lake Þingvallavatn and in streams connected to the lake. The research questions were,
1) Where in the lake and connected streams are juvenile fish found?
2) Has the density of charr and trout juveniles changed in the last twenty years?
3) Is there a connection between the environmental conditions and the presence of juvenile fish?
Data from Veiðimálastofnun (now Hafrannsóknastofun) surveys of salmonid juveniles in Þingvallavatn and adjacent rivers, spanning 2000 to 2021, were analyzed. In the summer of 2022, ten locations in Þingvallavatn were surveyed, fish were caught and measured, and several environmental factors were assessed using electrofishing. Fish were counted, identified by species, and measured for length and weight. Additionally, environmental factors, including the lake bottom composition and shoreline vegetation, were documented. The results show that the two species utilize different juvenile habitats, trout predominantly occupy river habitats, whereas charr are more abundant in the surf zone of the lake. There was a significant increase in density of the Brown trout over the 19-year period, but density of Arctic charr remained relatively stable with a sign of subtle decline. In 2022, six possible nursery sites were documented in the surf zone, four with majority of Arctic charr and two with mainly Brown trout. There was little overlap in the species distribution. Vegetation on the shoreline was the only environmental factor significantly correlated with the presence or absence of juvenile fish on a given site. Further studies on juvenile fish biology in the lake and adjacent streams and river might explore the interplay of environmental factors and genetics in shaping the size and morphology of adult fish in both species´ populations. This is particularly relevant for Arctic charr, which features four distinct sub-populations (or morphs) differing in size, shape, trophic morphology and ecology.
Mývatn conference
Lake Mývatn in north Iceland is unique for its biota and geology. To commemorate 50 years of research on the lake, and the retirement of Árni Einarsson a meeting was held there last week. Multiple scientists came to describe the hydrology, ecology and evolution of the organisms and ecosystem in the lake.
The programme was quite extensive and interesting - Mývatn research conference 2024.
Guðbjörg and Kenedy brought some of our fish stuff to the meeting.
Guðbjörg Ósk Jónsdóttir - Divergence of functional phenotypes in feeding elements of Arctic charr along the benthic – pelagic axis
Kenedy A. Williams - Patterns of Feeding Element Specialization in Arctic charr and Brown trout
Arnar Pálsson - Genetic structure and relatedness of anadromous and landlocked populations of Icelandic brown trout